UCLUST sort order
See also
UCLUST algorithm
Abundance sort
Sort order
UCLUST assumes that input sequences are sorted in an order such that an appropriate centroid sequence is found before other members of its cluster. The two most common sort orders are summarized in the table below.
|
Order |
Description |
|
Decreasing length |
This order is most appropriate when both full-length sequences and fragments are present, as shown in the figure below. However, with a length sort, the longest sequence may be an outlier. |
|
Decreasing abundance |
See abundance sorting . |
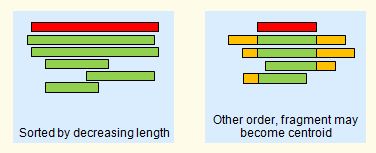
Multiple alignment of a cluster.
The centroid (representative) sequence is shown in red.
Fragments are poor centroids because member sequences may be

