UCHIME and DECIPHER
See also
Chimera benchmark home
Validation of DECIPHER on published benchmarks
DECIPHER is a 16S chimera detection method published in 2012. The authors claim that DECIPHER has performance comparable to UCHIME, especially with longer sequences and when the parents of the chimera are at least 10% diverged from each other. These claims are based mostly on results measured on their own simulated chimeras. Method rankings on previously published benchmarks are not reported in the paper. My results show that DECIPHER in fact has substantially lower sensitivity than UCHIME, even in those regimes where the authors claim DECIPHER has highest accuracy. It is not clear to me why the performance of DECIPHER is better on the authors' own test set; overtraining is one possible explanation.
Benchmark sets
I tested DECIPHER on the two of the benchmarks we used in the UCHIME paper: SIM2 and MOCK, both of which were developed by authors of previously published chimera methods. SIM2 is a set of simulated chimeras from the Haas et. al ChimeraSlayer paper. SIM2 has bimeras from 200nt to full-length genes with zero to 5% added noise. MOCK contains denoised reads from several mock community experiments in the Quince et al. in the Perseus paper. These reads are inferred to be "good" (i.e., non-chimeric) or to be chimeras by mapping reads to the known 16S sequences of the individually sequenced type strain bacteria in these communities. Reads of chimeric amplicons in these experiments are observed to have two, three or four segments. The MOCK experiments used wet-bench sequencing in which chimeras form naturally as a side effect of amplification, avoiding the assumptions and simplifications required for in silico simulation.
Results
On all sets I've tested so far, the sensitivity of DECIPHER is substantially lower than UCHIME, as shown in the table and figure below. I tried to identify subsets that would support the authors' claim that DECIPHER works best with longer sequences and when parent divergence is high, but even in these cases DECIPHER has poor performance.
|
Benchmark and subset
|
UCHIME
sensitivity
|
DECIPHER
sensitivity
|
|
MOCK Uneven, all chimeras
|
94% |
63% |
|
MOCK Uneven, div >= 10%
|
89% |
77% |
| SIM2, full-length, 2% indel noise, D >= 20% |
100% |
69% |
| SIM2, L=300, no mutations, D >= 10% |
99% |
46% |
| SIM2, L=300, no mutations, all chimeras |
81% |
31% |
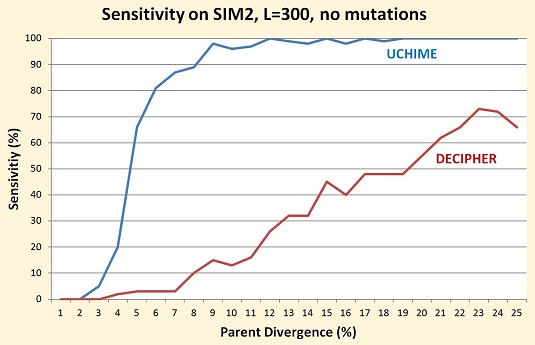
Comment on Table 1 in the DECIPHER paper
In Table 1, the authors present DECIPHER results for SIM2 sets with from 1 to 5% mutations. Only chimeras with at least 20% parent divergence were included (the motivation for this choice is not explained), and UCHIME results are not included. DECIPHER is reported to have from 31% to 76% sensitivity, while UCHIME has 100% sensitivity on all subsets.