See also
UPARSE home page
UPARSE pipeline
OTU benchmark methods and data
Introduction
In SSU metagenomics,
next-generation reads are clustered into
Operational Taxonomic Units (OTUs). This requires
quality filtering,
dereplication,
discarding singletons (optional), and finally
clustering into OTUs, typically at a 97% identity threshold.
Benchmark results
The OTU benchmark uses 454 Titanium
and Illumina MiSeq reads of Even and Staggered mock communities used for
protocol development in the Human
Microbiome Project (HMP). USEARCH results were obtained with the same parameters for all samples.
The number of reads per sample ranges from 10,000 (Titanium) to two million (MiSeq). The
accuracy of UPARSE was compared to
recommended procedures (Sept. 2012) for mothur, QIIME and AmpliconNoise.
| Accuracy
measure |
Summary |
Detailed results
(click on image) |
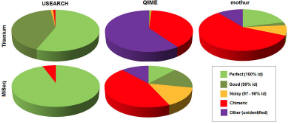
Sequence quality
Are OTUs accurate reconstructions of biological sequences? |
Most USEARCH OTUs are >=99% identical to a biological
sequence. Most QIIME, mothur and AmpliconNoise OTUs are >3%
diverged from a biological sequence. Roughly half are chimeric. |
|
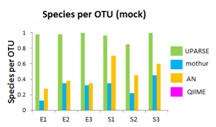
Diversity
Does the number of OTUs correspond to the number of
species? |
USEARCH generated from 0.8 to 1.0 OTUs per
detectable species. Mothur
and AmpliconNoised produced 2.3x to 6.7x more OTUs
than species.
QIIME produced thousands of OTUs, far more than the
number of species. |
|
Reference
Edgar, R.C. (2013) UPARSE: Highly accurate OTU sequences from microbial amplicon reads,
Nature Methods [Pubmed:23955772,
dx.doi.org/10.1038/nmeth.2604].
|


